Documentation
Input
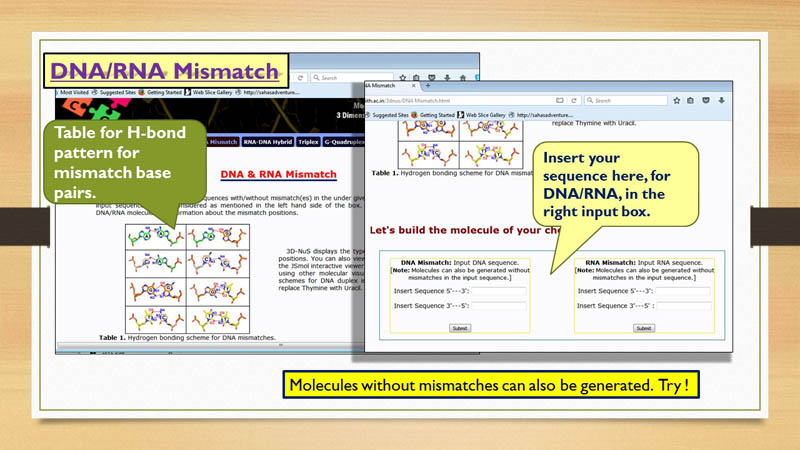
DNA/RNA Mismatch
3D-NuS offers three options for building either ideal models, sequence specific models or hairpins of RNA or DNA duplex
with or without mismatches. While the 'Ideal model' option builds the duplex using ideal B-DNA and A-RNA parameters, the 'sequence specific model' builds
the duplex using sequence specific base step and base pair parameters. Sequence specific parameters are extracted from the survey and 3DNA analysis of all NMR and
and X-Ray derived PDB structures of DNA and RNA (till the date 14-06-2017) and the average values are presented in Table S1 and S2 respectively with standard deviation presented in red color.
In analysis of PDB files, apo form of DNA and RNA double helical structures are only considered and non-Watson-Crick base pairs are excluded from analysis.
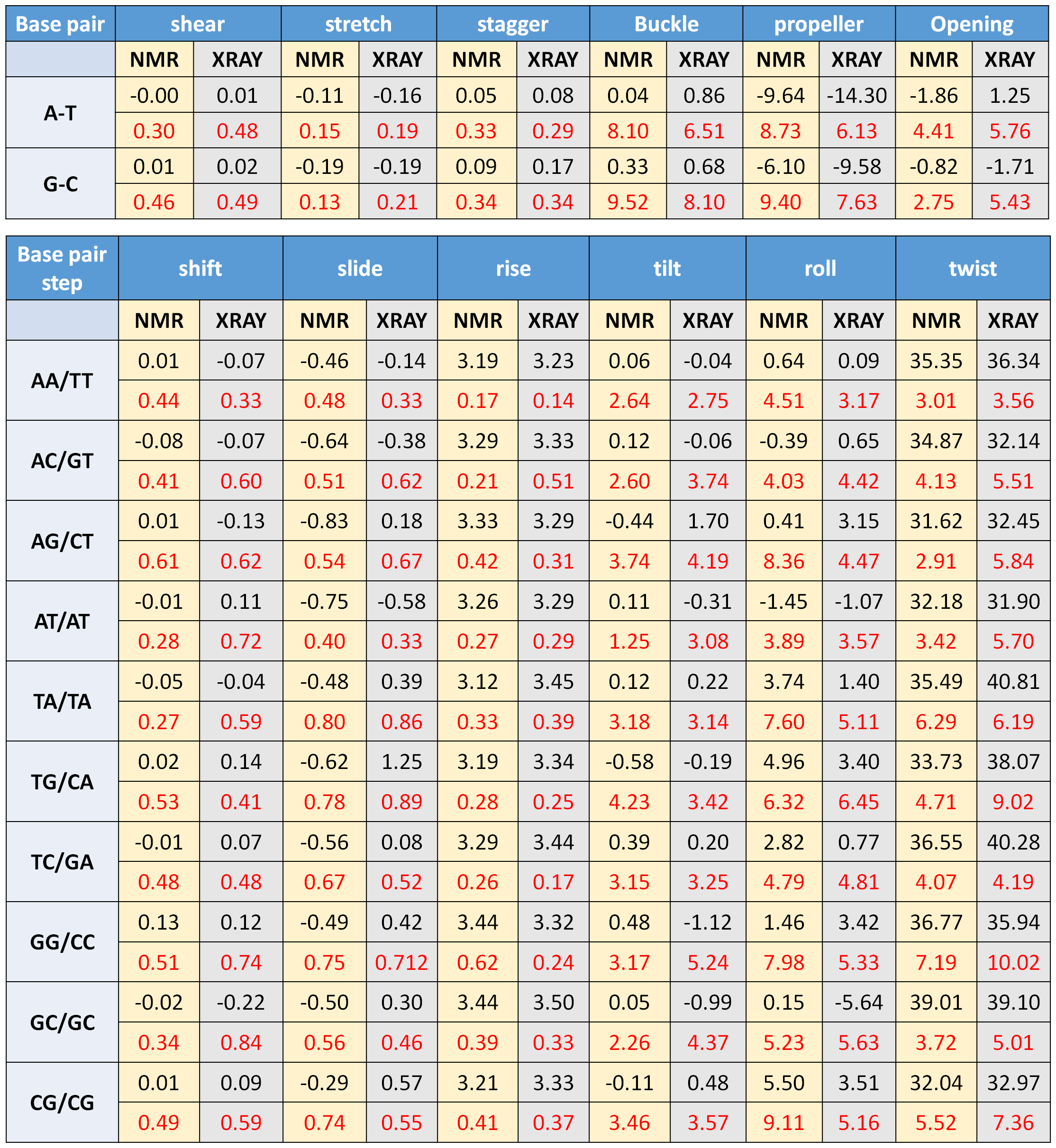
Table S1, © IIT Hyderabad
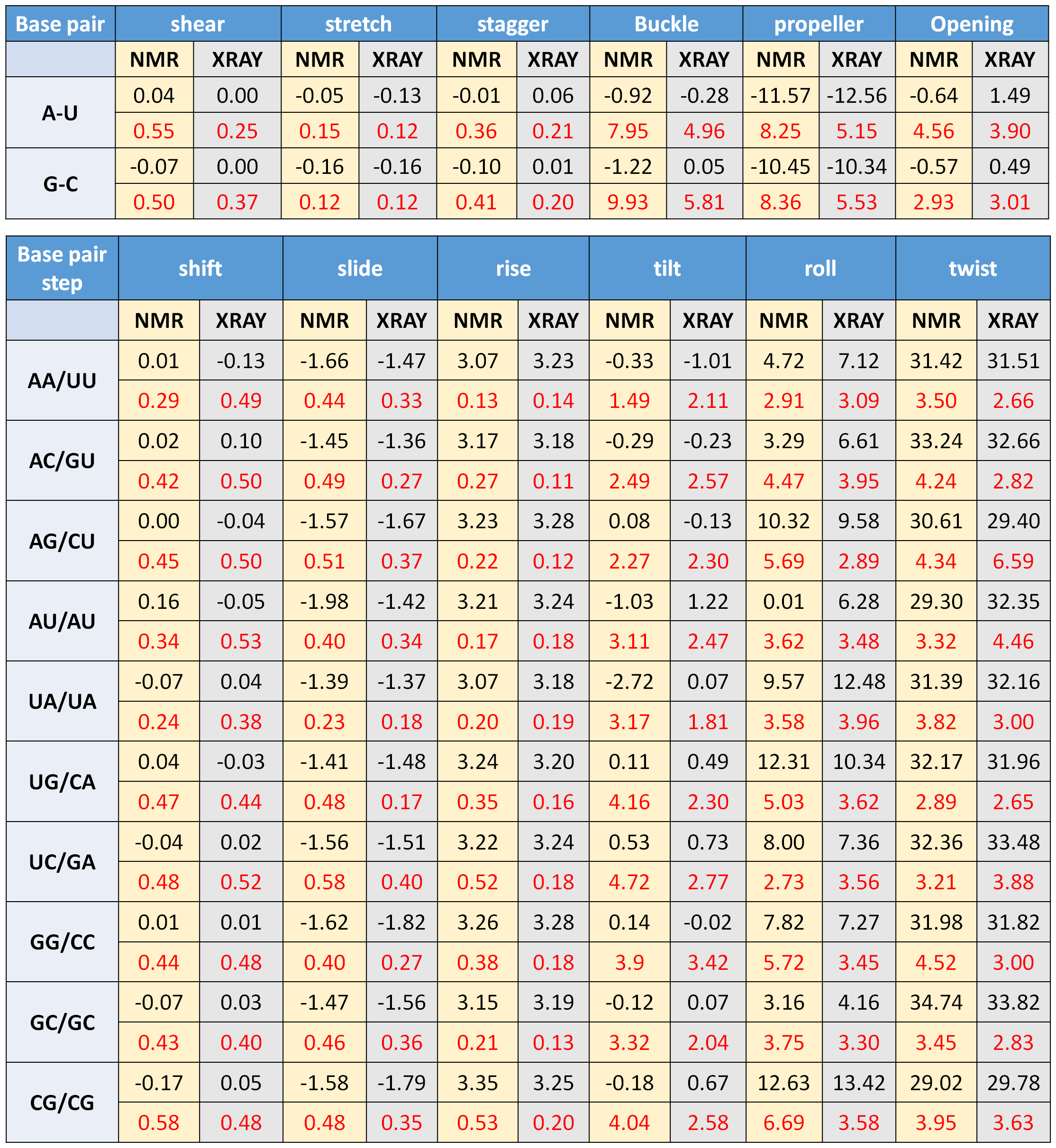
Table S2, © IIT Hyderabad
For generating DNA/RNA duplexes, user need to provide the
nucleic acid sequence for both the strands. Hydrogen bonding patterns in mismatch pairs are shown
in Figure 1 of DNA/RNA Mismatch page.
This information is useful in understaning the mode of
interaction in case of mismatch pairs. The orientation of the sequence for different strands
are mentioned in the input box. There is no upper limit for number of mismatch pairs or length of
the sequence. The conventional duplex structure parameters and aforementioned hydrogen bonding
pattern for mismatches are considered for building the duplexes.
Additionally, the server can also generate intramolecular DNA/RNA duplexes (hairpin structures)
for user defined loop sequences. Though the models are steric free, the conformations corresponding
to the loop residues may not be realistic. However, they can act as reasonably good starting structures for modeling studies.
DNA/RNA Chimera
3D-NuS also constructs chimeric structures, wherein, position specific ribo-/deoxy ribo- nucleotides can be introduced in the DNA and RNA duplexes respectively. User has to specify the template duplex sequences in DNA/RNA Chimera, followed by the choice of position and nucleotide to be introduced in the chimera.
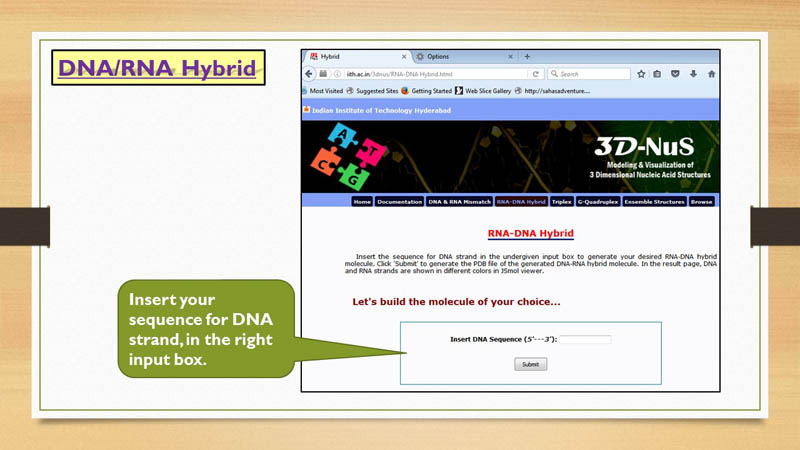
RNA-DNA Hybrid
3D-NuS allows two type of RNA-DNA hybrid models, "Ideal Model" and "Sequence specific model". While the 'Ideal model' option builds the duplex using ideal hybrid structural parameters, the 'sequence specific model' builds
the duplex using sequence specific base step and base pair parameters. Sequence specific parameters are extracted from the survey and 3DNA analysis of all NMR and
and X-Ray derived PDB structures of RNA-DNA hybrids (till the date 14-06-2017) and the average values are presented in Table S3 with standard deviation presented in red color.
In analysis of PDB files, apo form of RNA-DNA hybrid structures are only considered and chimera structures are excluded from analysis. As very few hybrid structures are available in PDB database, all base step and base pair parameters could not be fetched.
So, only available base pair and base step parameters are tabulated and for others ideal values are considered to build the models.
As 3D-NuS does not allow mismatches in RNA-DNA hybrid models, user need to give nucleic acid sequence for the DNA strand only and that should be in 5'-3' orientation.
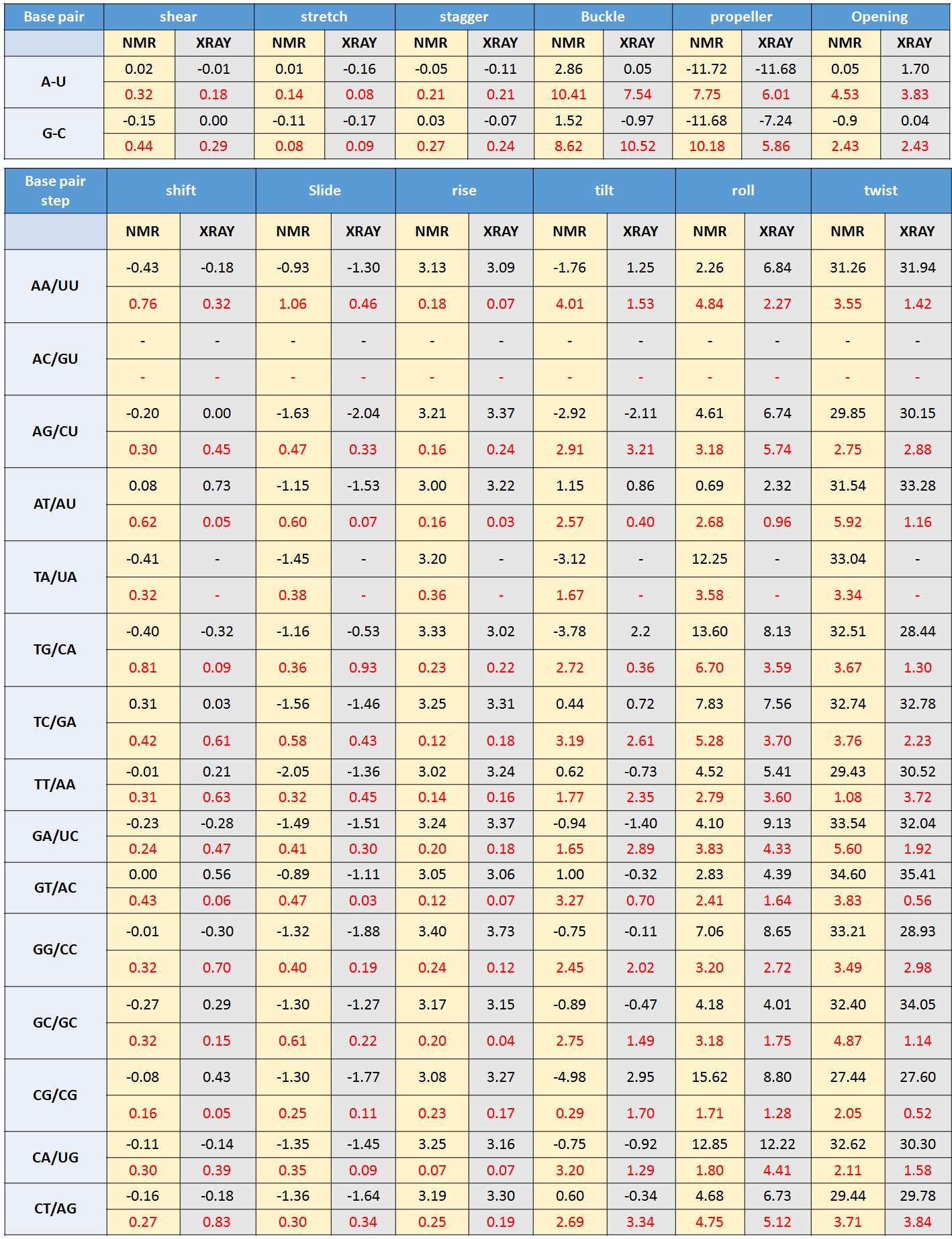
Table S3, © IIT Hyderabad
Density plots for base pair and base step parameters of DNA, RNA and RNA-DNA duplex.
All B-DNA, A-RNA and DNA-RNA hybrid structures deposited in PDB server are analyzed to derive base pair and base pair step parameters using the method described in Patro et. al (Accepted for publication in JMB). Density plots were made corresponding to these base pair and base pair step parameters. Go to this LINK for all density plots and further details.
Z-DNA/RNA
3D-NuS builds Z-DNA and Z-RNA duplexes by considering structural parameters corresponding to ideal Z-form geometry. As (CG)n rich sequences are prone to adapt Z-form, the user has to simply insert the number of CG repeats to generate the Z-form duplexes of DNA and RNA using 3D-NuS.
Triplex
To our knowledge, 3D-NuS is the first web-server that provides a provision
for modeling nucleic acid triplexes of 80 different types. Triplexes are purine/pyrimidine
rich homomeric repeat structures, wherein, the third strand mostly interacts with purine
rich strand of the duplex. Only few combinations of nucleotides are favorable in the third
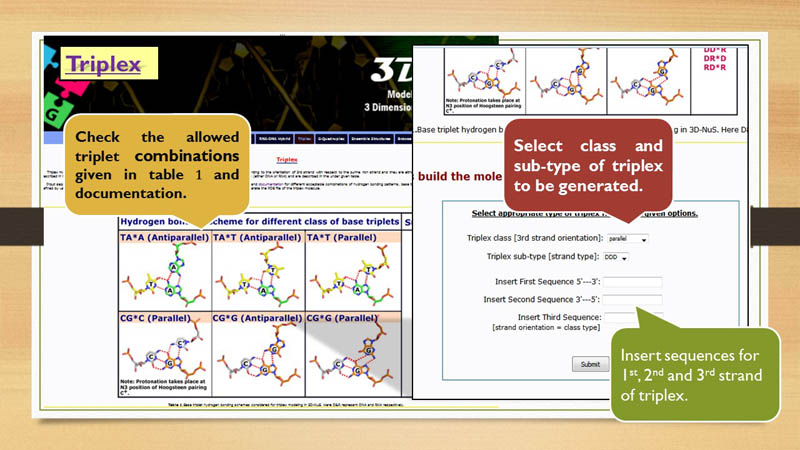
strand, based on the experimentally observed stability. Possible combinations of triplets
and the third strand orientations are described under Figure 2 of Triplex modeling
page. In total, 2 classes and 8 subtypes of triplexes can be built through 3D-NuS server.
The classes are defined with repect to orientation of third strand to the duplex, either
parallel (parallel triplex formed by Hoogsteen hydrogen bonds) or anti-parallel (antiparallel
triplex formed by reverse Hoogsteen hydrogen bonds) (copy from the manuscript). Similarly,
8 subtypes are defined with respect to the nature of individual strands i.e. DNA or RNA and
their different combinations. User need to select these option using the drop down to generate a particular triplex model.
Note that the user needs to insert the sequences for all three strands. Input
sequences for first and second strand must be of pyrimidine and purine nucleotides respectively
and the third strand can be of either of purine nucleotides or pyrimidine nucleotides or
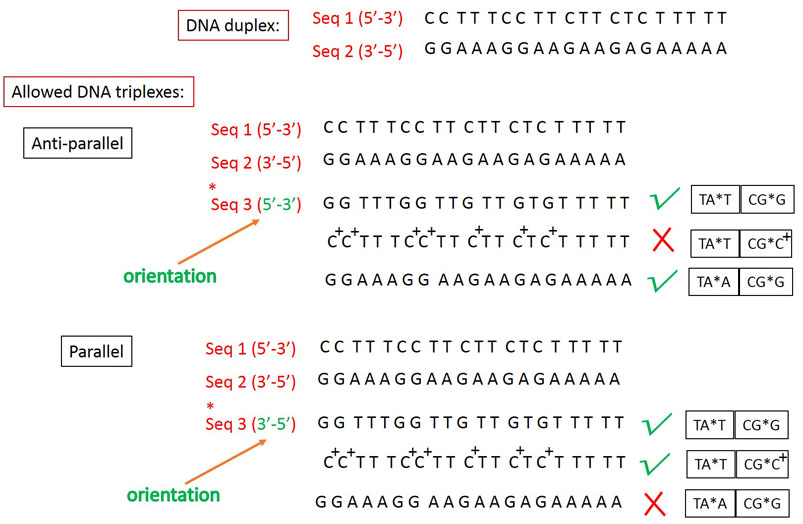
combination of both. But, it is very important to remember that only particular combination of
triplets are allowed as described in the example below. Taking these into consideration, the
user can built 80 different triplex models.
Example:
G-Quadruplex
3D-NuS can generate both inter- and intra- molecular G-quadruplex structures.
As a detailed protein data bank (PDB)-wide survey has indicated 58 possible folds that a DNA/RNA G-quadruplex can adapt,
the server generates 17 classes of G-quadruplex folds (Qn, wherein n=1 to 17). Among them, Q1 to Q5 (frequently found)
and Q14 to Q17 (rarely found) are monomeric folds, while Q6 to Q9 are dimeric folds. In contrast, Q10 to Q13 are intermolecular
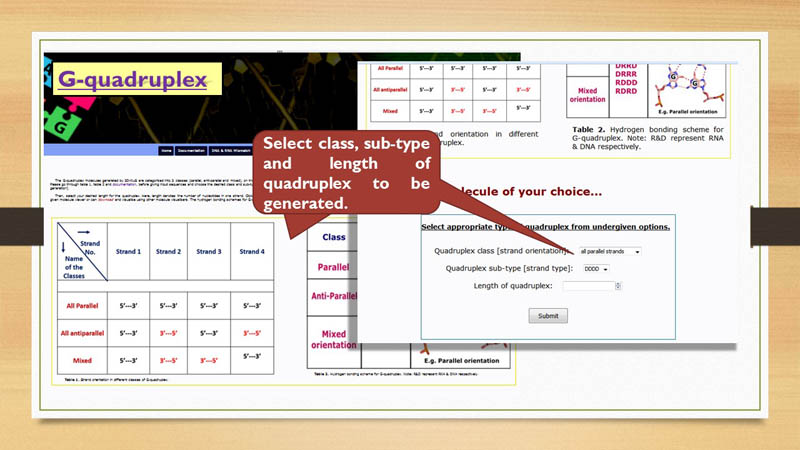
(tetrameric) in nature. For each class, a few sub-classes are supported on the basis of the molecular nature (DNA or RNA).
For instance, two sub-classes are supported for the monomer: either DNA (D) or RNA (R).
Four sub-classes are possible for the dimer: DD, RR, RD and DR. Tetramer can have 6 sub-classes: DDDD, RRRR, DRRD, DRRR, RDDD and RDRD.
Thus, 3D-NuS can generate 58 varieties of G-quadruplexes. The user can select the choice of their quadruplex class and sub-class from
the dropdown menu along with the desired quaduplex length. The four guanines involved G-quadrate formation are interacting with each
other through Hoogsteen hydrogen bonding scheme accompanied by syn or anti glycosyl conformation based on the orientations of the guanine
strands. The guanines are in anti conformation, when two guanine strands are parallel to each other (Qn, wherein n=2-5, 7-10, 12-17).
However, when two guanine strands are antiparallel to each other as in Q1, Q2, Q4-Q7, Q9, and Q11-Q17, they are related by syn and anti
glycosyl conformations. For the monomeric and dimeric quadruplexes, the server gives freedom for the user to specify the loop sequences
of their preference. Subsequently, the server establishes the connectivity between the strands using the user defined sequence.
Nevertheless, the sugar-phosphate conformations of loop regions of intramolecular quadruplex structures are bound to a limitation that
they are generated so as to have a steric free model and may not represent the realistic situation.
User can generate different models of G-quadruplex by providing class, sub-class & length of one strand and inserting sequence for the loop.
Output
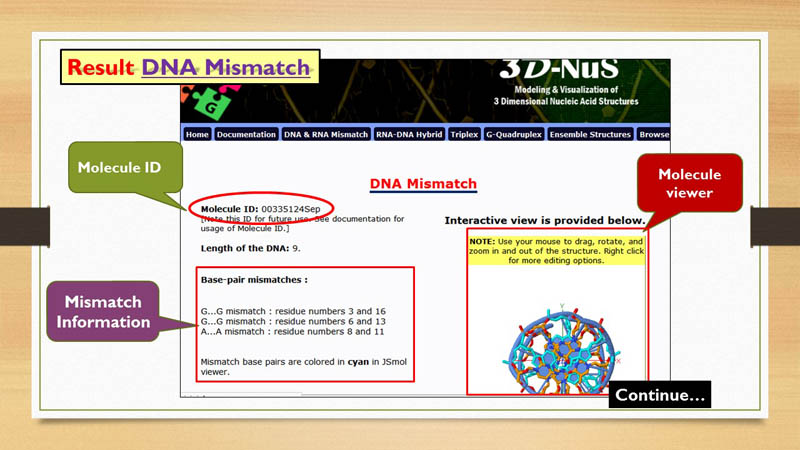
Molecule ID
Every nucleic acid model generated by 3D-NuS server, has an unique molecule ID. This molecule id can be used by the user to generate
ensemble of a particular structure and retrieve the same molecule later, after closing the session. This molecule ID expires in 7 days. This means,
a user can acess 3D-NuS generated nucleic acid molecule within a week only.
NoteMolecule ID must not be mis-understood as molecule name. Names of 3D-NuS generated molecules are in following format:
Name- "molecule type"_3dnus E.g.- rna_3dnus
And the same name is used for naming corresponding PDB file.
E.g.- rna_3dnus.pdb
Molecule ID is assigned based on the time and date of the job submitted.
E.g.- 02354724Sep
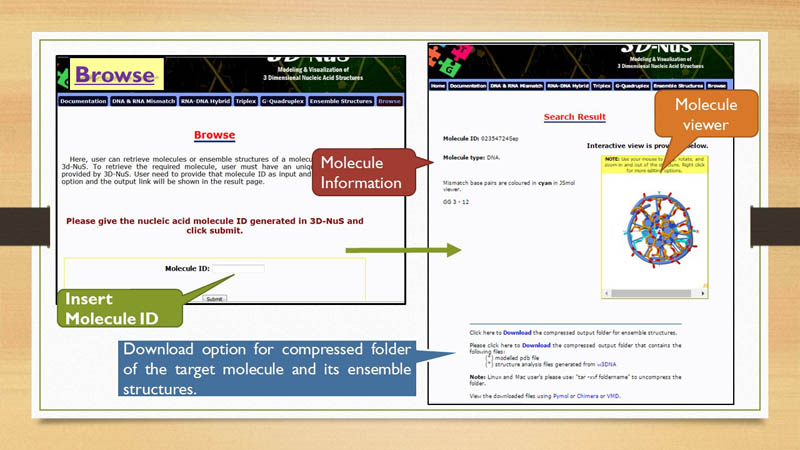
Molecule Viewer
3D-NuS server has embeded JSmol applet to view 3 dimensional structures of the generated nucleic acid molecule. This is an
interactive window for the user. User can rotate, zoom in, zoom out and change different representation of the molecule for better visualization.
Non-canonical base pairs of mismatches and strands like third strand in triplex, rna strand in RNA-DNA hybrid are highlighted
with different colors in JSmol viewer.
Nucleotide numbering pattern
3D-NuS numbers the nucleotides in the given strand from 5' end and continues in the same fashion for 2nd, 3rd and 4th strands as applicable. See below for explanations.
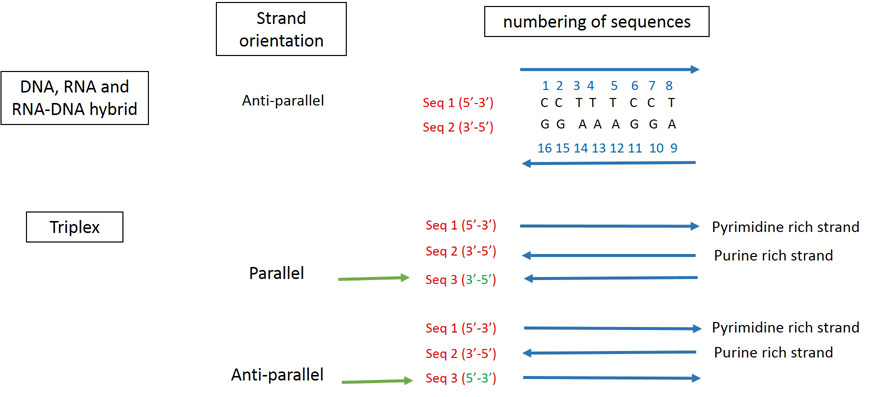
Mismatch Information
Mismatch base-pairs in DNA and RNA are shown in the result page with respective positions in the molecule. The residue numbers assigned to mismatch nucleotide bases are reffered from nucleotide numbering pattern explained in previous section. These mismatch base-pairs are represnted in cyan color in JSmol window.
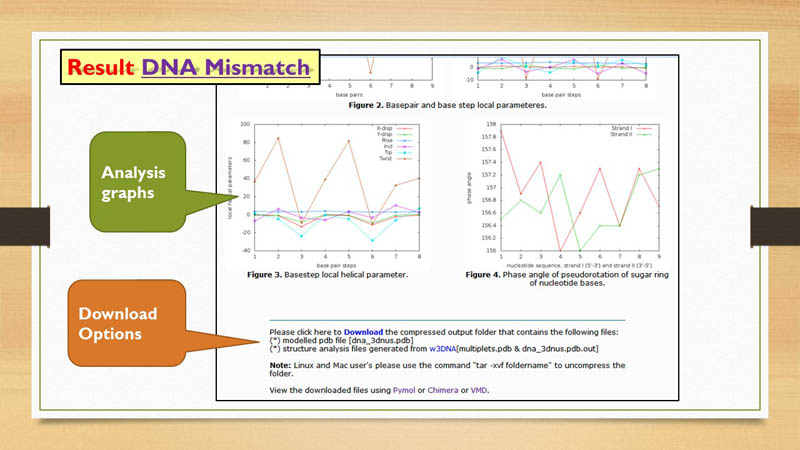
Structural Analysis
3D-NuS generated structures are further analysed by X3DNA software. Results are represented in graphs. Backbone torsion angles, base-pair, base-pair step & helical parameters, phase angle of pseudo rotation (sugar pucker) are shown in graphs with respect to residue number in X-axis. These graphs can also be downloaded using download option.
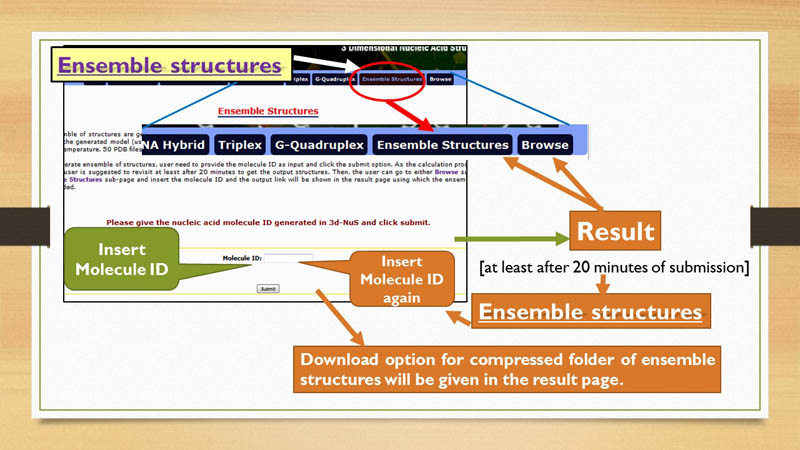
Ensemble Structures
Once a model is generated through 3D-NuS, a set of 50 models of same sequence with slightly different conformations can also be generated by 3D-NuS in Ensemble sub-page. For this, the user need to have unique molecule ID that is provided along with the modeled structure in the result sub-page.
Downloads
3D-NuS provides two types of downloading options. These download options refer to two compressed folders in "tar" format. When user generates a model for first time in 3D-NuS server, the first download option appears, which contains 3 dimensional coordinates of the generated molecule in PDB format and structural analysis files. The second download option is linked to the folder containing ensemble of structures, if user has chosen to generate the same. This option is shown after providing the molecule ID in Browse page only if ensemble structures are generated for that unique molecule ID.
Browse
Browse page in 3D-NuS provides an option to retrieve files generated through 3D-NuS server in past. The time limit for retrieving molecule structures is seven days. After that the folder containg information ragarding a particular molecule ID is erased from server and user cannot acess it any further.